from datascience import *
import numpy as np
import matplotlib
from mpl_toolkits.mplot3d import Axes3D
%matplotlib inline
import matplotlib.pyplot as plots
plots.style.use('fivethirtyeight')
import warnings
warnings.simplefilter("ignore")ckd = Table.read_table('ckd.csv')
ckd = ckd.relabeled('Blood Glucose Random', 'Glucose').select('Glucose', 'Hemoglobin', 'White Blood Cell Count', 'Class')patients = Table.read_table('breast-cancer.csv').drop('ID')
def randomize_column(a):
return a + np.random.normal(0.0, 0.09, size=len(a))
jittered = Table().with_columns([
'Bland Chromatin (jittered)',
randomize_column(patients.column('Bland Chromatin')),
'Single Epithelial Cell Size (jittered)',
randomize_column(patients.column('Single Epithelial Cell Size')),
'Class',
patients.column('Class')
])Google Science Fair¶
patients = Table.read_table('breast-cancer.csv').drop('ID')
patients.show(3)Loading...
patients.group('Class')Loading...
patients.scatter('Bland Chromatin', 'Single Epithelial Cell Size', group='Class')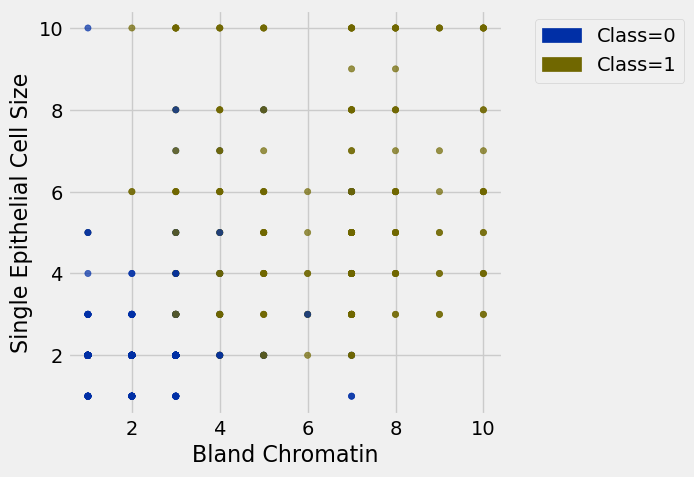
jittered.scatter(0, 1, group='Class')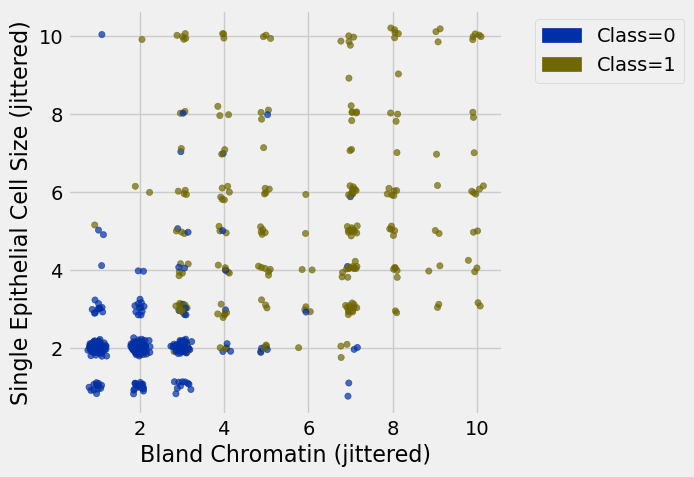
Distance¶
def distance(pt1, pt2):
"""Return the distance between two points, represented as arrays"""
return np.sqrt(sum((pt1 - pt2)**2))example1 = make_array(4, 2)
example2 = make_array(4, 3)
distance(example1, example2)1.0(example1 - example2)** 2array([0, 1])np.array(patients.drop('Class').row(0))array([5, 1, 1, 1, 2, 1, 3, 1, 1])def row_distance(row1, row2):
"""Return the distance between two numerical rows of a table"""
return distance(np.array(row1), np.array(row2))attributes = patients.drop('Class')
attributes.show(3)Loading...
row_distance(attributes.row(0), attributes.row(1))11.874342087037917row_distance(attributes.row(0), attributes.row(2))2.2360679774997898row_distance(attributes.row(2), attributes.row(2))0.0The Classifier¶
def distances(training, example):
"""
Compute distance between example and every row in training.
Return training augmented with Distance column
"""
distances = make_array()
attributes_only = training.drop('Class')
for row in attributes_only.rows:
distances = np.append(distances, row_distance(row, example))
# ^ SAME AS DOING:
#
# for i in np.arange(attributes_only.num_rows):
# row = attributes_only.row(i)
# distances = np.append(distances, row_distance(row, example))
return training.with_column('Distance_to_ex', distances)example = attributes.row(21)
exampleRow(Clump Thickness=10, Uniformity of Cell Size=5, Uniformity of Cell Shape=5, Marginal Adhesion=3, Single Epithelial Cell Size=6, Bare Nuclei=7, Bland Chromatin=7, Normal Nucleoli=10, Mitoses=1)distances(patients.exclude(21), example).sort('Distance_to_ex')Loading...
def closest(training, example, k):
"""
Return a table of the k closest neighbors to example
"""
return distances(training, example).sort('Distance_to_ex').take(np.arange(k))closest(patients.exclude(21), example, 5)Loading...
closest(patients.exclude(21), example, 5).group('Class').sort('count', descending=True) # .column('Class').item(0)Loading...
def majority_class(topk):
"""
Return the class with the highest count
"""
return topk.group('Class').sort('count', descending=True).column(0).item(0)def classify(training, example, k):
"""
Return the majority class among the
k nearest neighbors of example
"""
return majority_class(closest(training, example, k))classify(patients.exclude(21), example, 5)1patients.take(21)Loading...
new_example = attributes.row(10)
classify(patients.exclude(10), new_example, 5)0patients.take(10)Loading...
another_example = attributes.row(15)
classify(patients.exclude(15), another_example, 5)0patients.take(15)Loading...
Review of the Steps¶
distance(pt1, pt2): Returns the distance between the arrayspt1andpt2row_distance(row1, row2): Returns the distance between the rowsrow1androw2distances(training, example): Returns a table that istrainingwith an additional column'Distance'that contains the distance betweenexampleand each row oftrainingclosest(training, example, k): Returns a table of the rows corresponding to the k smallest distancesmajority_class(topk): Returns the majority class in the'Class'columnclassify(training, example, k): Returns the predicted class ofexamplebased on aknearest neighbors classifier using the historical sampletraining
Accuracy of a Classifier¶
patients.num_rows683shuffled = patients.sample(with_replacement=False) # Randomly permute the rows
training_set = shuffled.take(np.arange(342))
test_set = shuffled.take(np.arange(342, 683))def evaluate_accuracy(training, test, k):
"""Return the proportion of correctly classified examples
in the test set"""
test_attributes = test.drop('Class')
num_correct = 0
for i in np.arange(test.num_rows):
c = classify(training, test_attributes.row(i), k)
num_correct = num_correct + (c == test.column('Class').item(i))
return num_correct / test.num_rowsdef predictions(training, test, k):
evaluate_accuracy(training_set, test_set, 5)0.9648093841642229evaluate_accuracy(training_set, test_set, 3)0.9706744868035191evaluate_accuracy(training_set, test_set, 11)0.9560117302052786evaluate_accuracy(training_set, test_set, 1)0.9648093841642229Standardize if Necessary¶
ckd.show(5)Loading...
shuffled = ckd.sample(with_replacement=False)
training_set = shuffled.take(np.arange(74))
test_set = shuffled.take(np.arange(74, 148))evaluate_accuracy(training_set, test_set, 3)0.7702702702702703def standard_units(x):
return (x - np.average(x)) / np.std(x)ckd_new = ckd.select('Class').with_columns(
'Glucose_su', standard_units(ckd.column('Glucose')),
'Hemoglobin_su', standard_units(ckd.column('Hemoglobin')),
'WBC_su', standard_units(ckd.column('White Blood Cell Count'))
)ckd_new.show(3)Loading...
shuffled = ckd_new.sample(with_replacement=False)
training_set = shuffled.take(np.arange(74))
test_set = shuffled.take(np.arange(74, 148))evaluate_accuracy(training_set, test_set, 3)0.9594594594594594